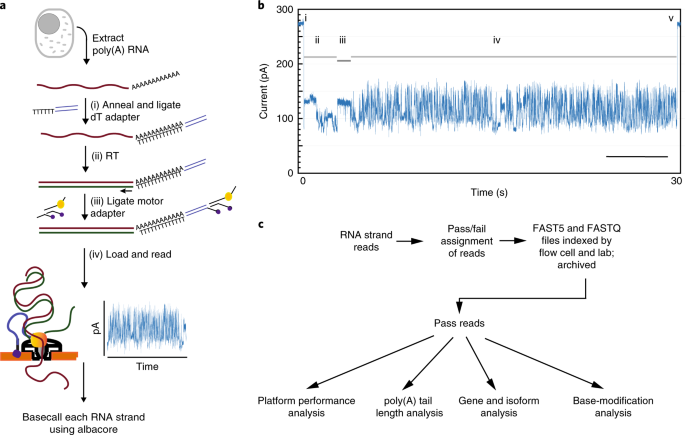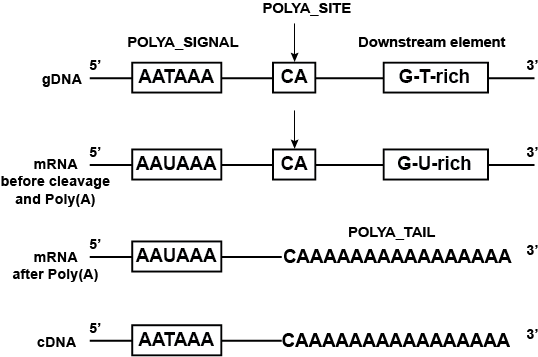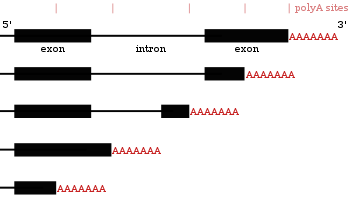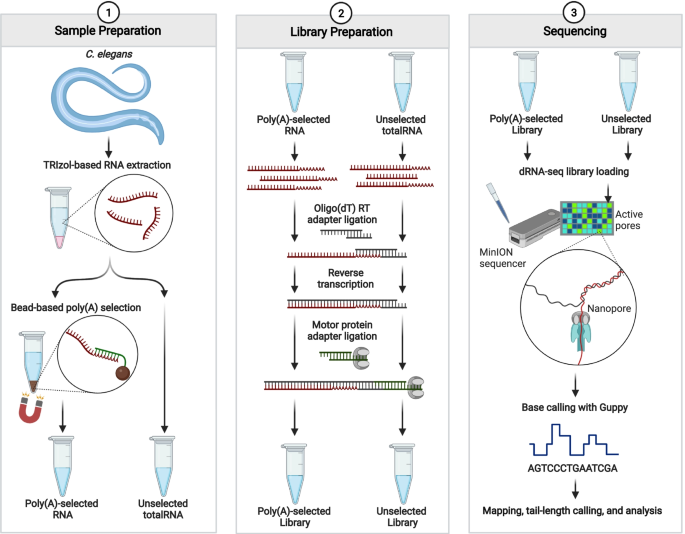
Poly(a) selection introduces bias and undue noise in direct RNA-sequencing | BMC Genomics | Full Text
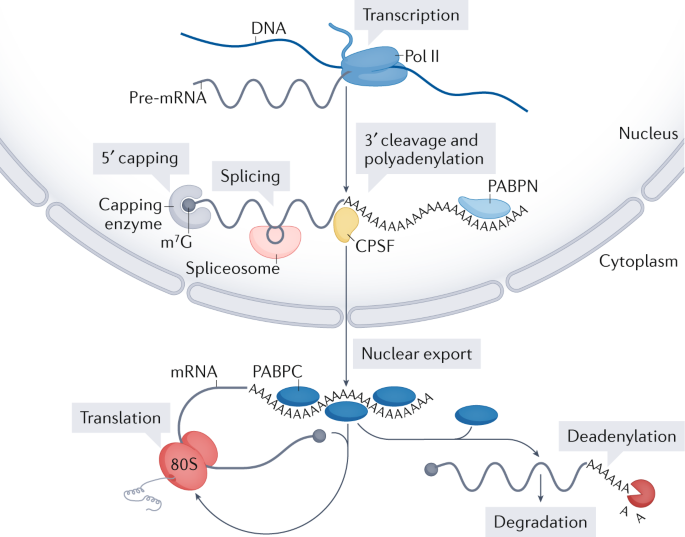
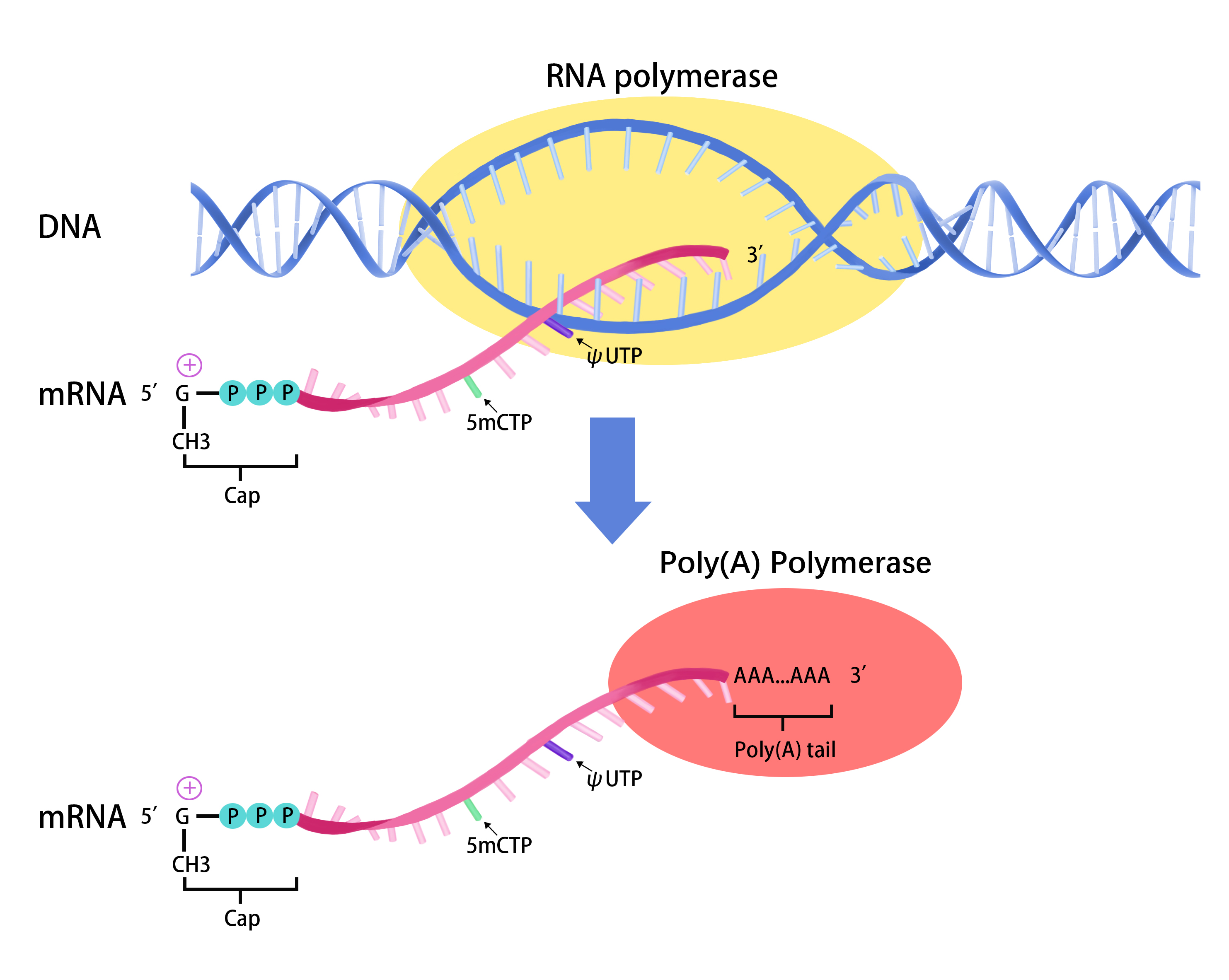
Roles of mRNA poly(A) tails in regulation of eukaryotic gene expression | Nature Reviews Molecular Cell Biology
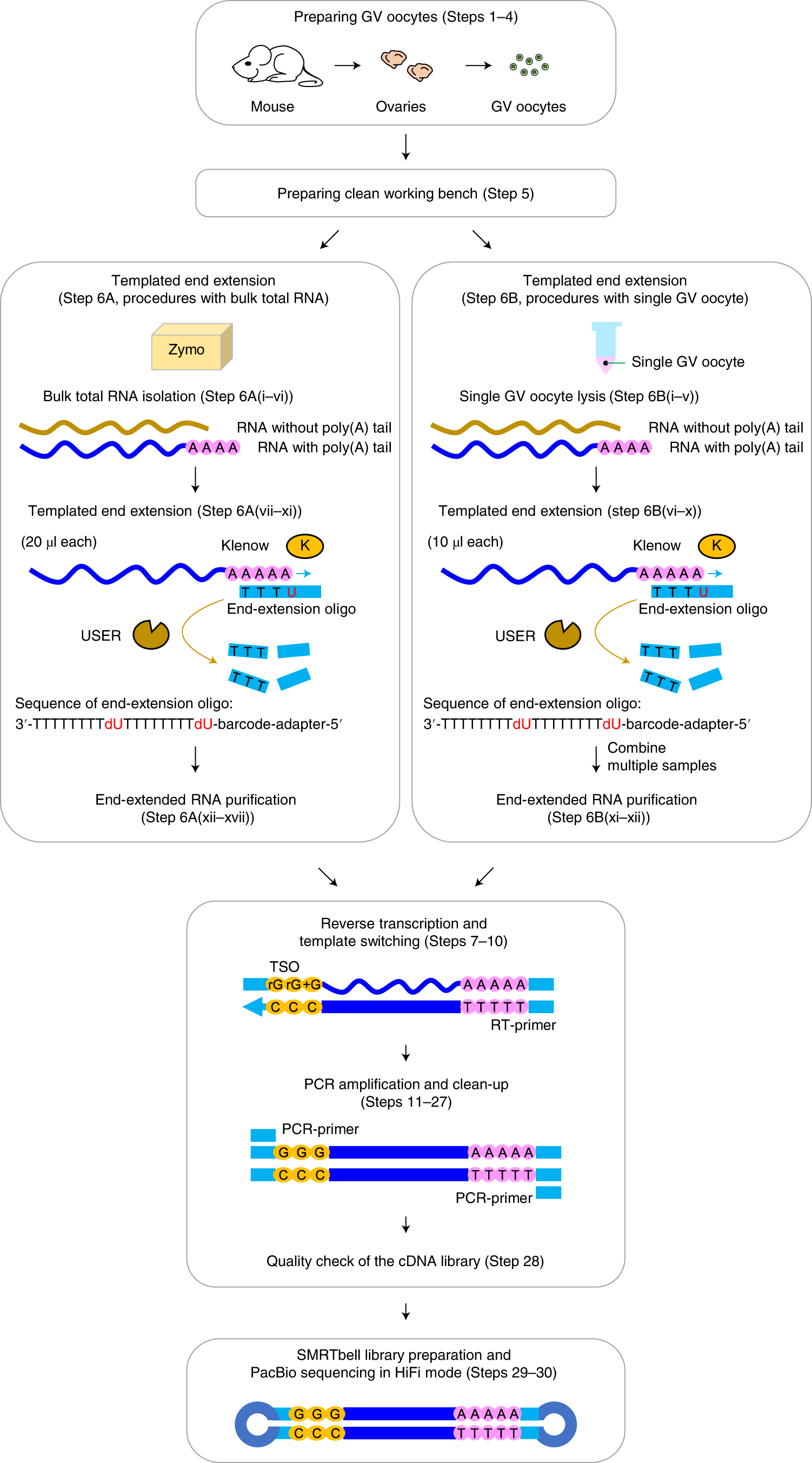
Transcriptome-wide measurement of poly(A) tail length and composition at subnanogram total RNA sensitivity by PAIso-seq | Nature Protocols

Transcription Dynamics Regulate Poly(A) Tails and Expression of the RNA Degradation Machinery to Balance mRNA Levels - ScienceDirect

RNA stabilization by a poly(A) tail 3′-end binding pocket and other modes of poly(A)-RNA interaction | Science
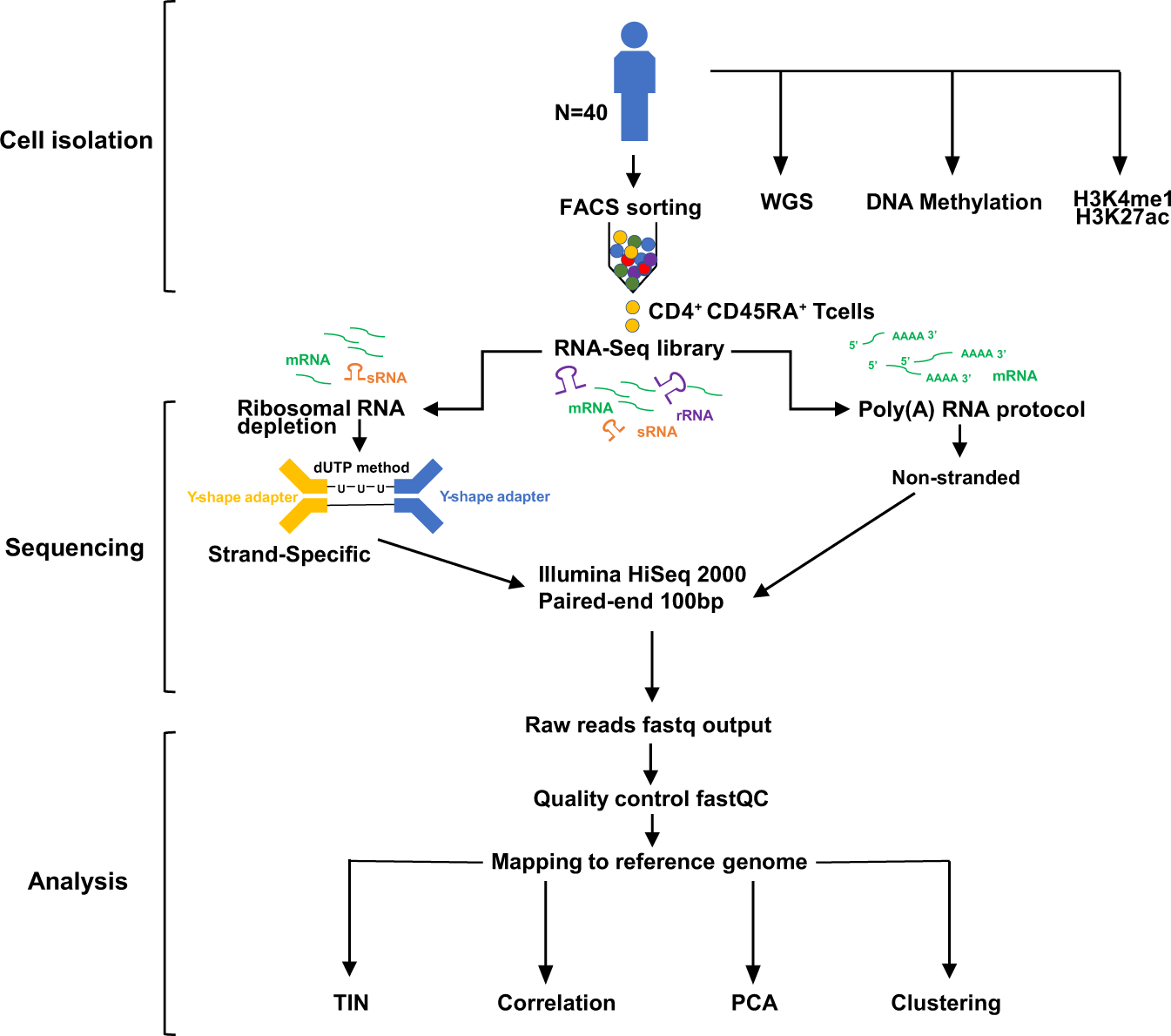
Paired rRNA-depleted and polyA-selected RNA sequencing data and supporting multi-omics data from human T cells | Scientific Data

The intrinsic structure of poly(A) RNA determines the specificity of Pan2 and Caf1 deadenylases | Nature Structural & Molecular Biology

RPAD (RNase R treatment, polyadenylation, and poly(A)+ RNA depletion) method to isolate highly pure circular RNA. - Abstract - Europe PMC

Poly(A)-seq – A method for direct sequencing and analysis of the transcriptomic poly(A)-tails | RNA-Seq Blog

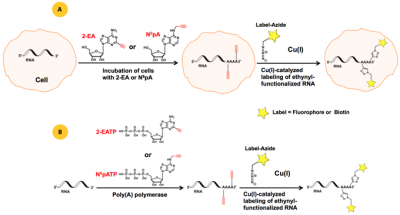
Poly(A)-ClickSeq – click-chemistry for next-generation 3΄-end sequencing without RNA enrichment or fragmentation | RNA-Seq Blog